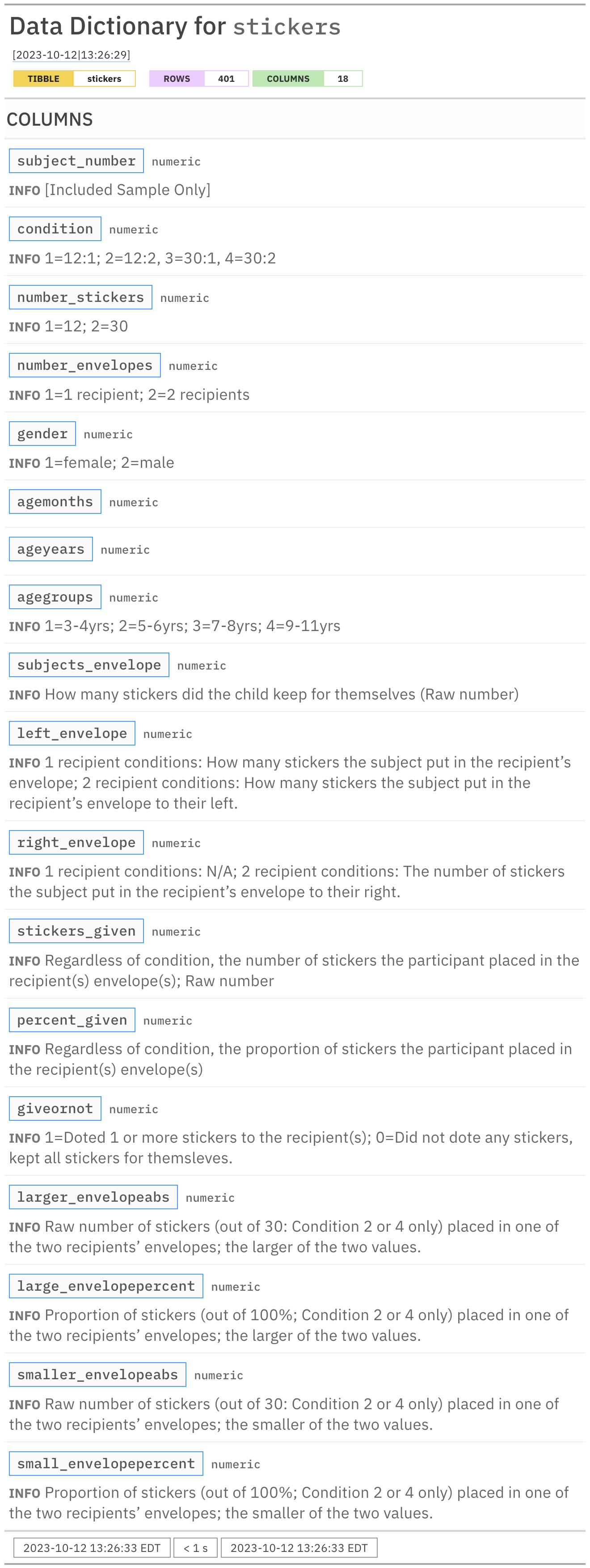
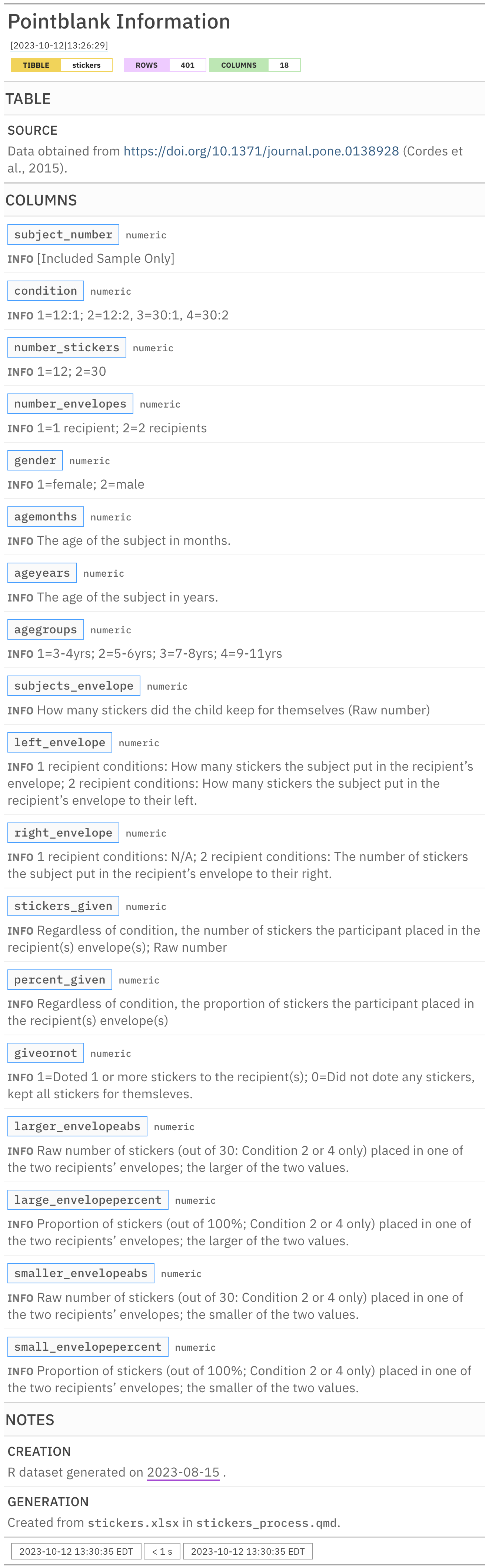
Rows: 401
Columns: 18
$ subject_number <dbl> 1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12, 13, 14, 1…
$ condition <dbl> 1, 1, 1, 1, 1, 1, 2, 2, 3, 3, 3, 3, 3, 4, 4, 4, …
$ number_stickers <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 2, 2, 2, 2, 2, 2, 2, 2, …
$ number_envelopes <dbl> 1, 1, 1, 1, 1, 1, 2, 2, 1, 1, 1, 1, 1, 2, 2, 2, …
$ gender <dbl> 1, 2, 2, 1, 2, 2, 1, 2, 2, 2, 2, 1, 1, 1, 2, 1, …
$ agemonths <dbl> 36, 36, 36, 36, 36, 36, 36, 36, 36, 36, 36, 36, …
$ ageyears <dbl> 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, …
$ agegroups <dbl> 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, 1, …
$ subjects_envelope <dbl> 7, 12, 4, 7, 12, 8, 8, 11, 26, 30, 12, 12, 30, 1…
$ left_envelope <dbl> 5, 0, 8, 5, 0, 4, 2, 1, 4, 0, 18, 18, 0, 11, 6, …
$ right_envelope <dbl> NA, NA, NA, NA, NA, NA, 2, 0, NA, NA, NA, NA, NA…
$ stickers_given <dbl> 5, 0, 8, 5, 0, 4, 4, 1, 4, 0, 18, 18, 0, 16, 7, …
$ percent_given <dbl> 0.42, 0.00, 0.67, 0.42, 0.00, 0.33, 0.33, 0.08, …
$ giveornot <dbl> 1, 0, 1, 1, 0, 1, 1, 1, 1, 0, 1, 1, 0, 1, 1, 1, …
$ larger_envelopeabs <dbl> NA, NA, NA, NA, NA, NA, 2, 1, NA, NA, NA, NA, NA…
$ large_envelopepercent <dbl> NA, NA, NA, NA, NA, NA, 0.5000000, 1.0000000, NA…
$ smaller_envelopeabs <dbl> NA, NA, NA, NA, NA, NA, 2, 0, NA, NA, NA, NA, NA…
$ small_envelopepercent <dbl> NA, NA, NA, NA, NA, NA, 0.5000000, 0.0000000, NA…